Publications
- Recalcitrant deep and shallow nodes in Aristolochia (Aristolochiaceae) illuminated using anchored hybrid enrichment. Wanke, S., Granados Mendoza, C., Müller, S., Paizanni Guillén, A., Neinhuis, C., Lemmon, A.R., Lemmon, E.M., Samain, M.S. (2017). Mol Phylogenet Evol. 2017 May 20. pii: S1055-7903(17)30366-4.
- The stability of the transcriptome during the estrous cycle in four regions of the mouse brain. DiCarlo, L.M., Vied, C., Nowakowski, R.S. (2017). J Comp Neurol. 525(15), 3360-3387.
- The phylogeny and biogeography of Hakea (Proteaceae) reveals the role of biome shifts in a continental plant radiation. Cardillo, M., Weston, P.H., Reynolds, Z.K.M., Olde, P.M., Mast, A.R., Lemmon, E.M., Lemmon, A.R., Bromham, L. (2017). Evolution. 71(8), 1928-1943.
- Proteomic characterization of paired non-malignant and malignant African-American prostate epithelial cell lines distinguishes them by structural proteins. Myers, J.S., Vallega, K.A., White, J., Yu, K., Yates, C.C., Sang, Q.A. (2017). BMC Cancer. 17(1), 480.
- Resolving relationships among the megadiverse butterflies and moths with a novel pipeline for Anchored Phylogenomics. Breinholt, J.W., Earl, C., Lemmon, A.R., Lemmon, E.M., Xiao, L., Kawahara, A.Y. (2017). Syst Biol. 2017 May 4.
- A pilot study applying the plant Anchored Hybrid Enrichment method to New World sages (Salvia subgenus Calosphace; Lamiaceae). Fragoso-Martínez, I., Salazar, G.A., Martínez-Gordillo, M., Magallón, S., Sánchez-Reyes, L., Lemmon, E.M., Lemmon, A.R., Sazatornil, F., Granados Mendoza, C. (2017). Mol Phylogenet Evol. 2017 Feb 9. pii: S1055-7903(16)30325-6.
- Resolving Rapid Radiations Within Angiosperm Families Using Anchored Phylogenomics. Léveillé-Bourret, É., Starr, J.R., Ford, B.A., Lemmon, E.M., Lemmon, A.R. (2017). Syst Biol. 2017 May 4.
- Sex differences in the molecular signature of the developing mouse hippocampus. Bundy, J.L., Vied, C., Nowakowski, R.S. (2017). BMC Genomics. 18(1), 237.
- Selection To Increase Expression, Not Sequence Diversity, Precedes Gene Family Origin and Expansion in Rattlesnake Venom. Margres, M.J., Bigelow, A.T., Lemmon, E.M., Lemmon, A.R., Rokyta, D.R. (2017). Genetics 206(3), 1569-1580.
- Stability of patient-specific features of altered DNA replication timing in xenografts of primary human acute lymphoblastic leukemia. Sasaki T, Rivera-Mulia JC, Vera D, Zimmerman J, Das S, Padget M, Nakamichi N, Chang BH, Tyner J, Druker BJ, Weng AP, Civin CI, Eaves CJ, Gilbert DM. (2017). Exp Hematol. 51, 71-82.
- In the shadows: Phylogenomics and coalescent species Delimitation Unveil Cryptic Diversity in a Cerrado Endemic Lizard (Squamata: Tropidurus). Domingos, F. M., Colli, G. R., Lemmon, A., Lemmon E. C., Beheregaray, L. B. (2017). Mol Phylogenet Evol. 107, 455-465.
- Neutral and selective drivers of colour evolution in a widespread Australian passerine. Morales, H. E., A. Pavlova, P. Sunnucks, R. Major, N. Amos, A. Wang, L. Joseph, A. R. Lemmon, J. A. Endler, K. Delhey. (2017). Journal of Biogeography. 44, 522-536.
- Human Mesenchymal Stem Cell Treatment Normalizes Cortical Gene Expression after Traumatic Brain Injury. Darkazalli, A., Vied, C., Badger, C.-D., & Levenson, C. W. (2017). Journal of Neurotrauma. 34(1), 204-212.
- Using phylogenomics to understand the link between biogeographic origins and regional diversification in ratsnakes. Chen, X., Lemmon, A.R., Lemmon, E.M., Pyron, R.A. Burbrink, F.T. (2017). Mol Phylogenet Evol. 111, 206-218.
- Anchored phylogenomics improves the resolution of evolutionary relationships in the rapid radiation of Protea L. Mitchell, N., Lewis, P.O., Lemmon, E.M., Lemmon, A.R., Holsinger, K.E. (2017). Am J Bot. 104(1), 102-115.
- Upregulation of minichromosome maintenance complex component 3 during epithelial-to-mesenchymal transition in human prostate cancer. Stewart, P.A., Khamis, Z.I., Zhau, H.E., Duan, P., Li, Q., Chung, L.W.K., Sang, Q.A. (2017). Oncotarget. 8(24), 39209-39217.
- Proteomic profiling of NCI-60 extracellular vesicles uncovers common protein cargo and cancer type-specific biomarkers. Hurwitz, S. N., Rider, M. A., Bundy, J. L., Liu, X., Singh, R. K., & Meckes, D. G. Jr. (2016). Oncotarget. 7(52), 86999-87015.
- Resolving Cypriniformes relationships using an anchored enrichment approach. Stout, C.C., Tan, M., Lemmon, A.R., Lemmon, E.M., Armbruster, J.W. (2016). BMC Evol Biol. 16(1), 244.
- Transcriptomic analysis of the hippocampus from six inbred strains of mice suggests basis for sex-specific susceptibility and severity of neurological disorders. Vied, C., Ray, S., Badger, C.-D., Bundy, J. L., Arbeitman, M. N., & Nowakowski, R. S. (2016). The Journal of Comparative Neurology. 524(13), 2696-7.
- Sex Differences in Drosophila Somatic Gene Expression: Variation and Regulation by Doublesex. Arbeitman, M. N., New, F., Fear, J. M., Howard, T. S., Dalton, J. E., & Graze, R. M. (2016). G3 (Bethesda, Md.). 6(7), 1799-808.
- Proteomic Upregulation of Fatty Acid Synthase and Fatty Acid Binding Protein 5 and Identification of Cancer- and Race-Specific Pathway Associations in Human Prostate Cancer Tissues. Myers, J.S., von Lersner, A.K., Sang, Q.X. (2016). J. Cancer. 7(11), 1452-1464.
- Diversification in wild populations of the model organism Anolis carolinensis: A genome-wide phylogeographic investigation. Manthey, J.D., Tollis, M., Lemmon, A.R., Lemmon, E.M., Boissinot, S. (2016). Ecol Evol. 6(22), 8115-8125.
- Fractionation-Dependent improvements in proteome resolution in the mouse hippocampus by IEF LC-MS/MS. Bundy, J. L., Inouye, B. D., Mercer, R. S., & Nowakowski, R. S. (2016). Electrophoresis. 37(14), 2054-62.
- Neurons that Underlie Drosophila melanogaster Reproductive Behaviors: Detection of a Large Male-Bias in Gene Expression in Fruitless-Expressing Neurons. Newell, N. R., New, F. N., Dalton, J. E., McIntyre, L. M., & Arbeitman, M. N. (2016). G3 (Bethesda, Md.). 6(8), 2455-65.
- Anchored enrichment dataset produced for true flies (order Diptera) reveals insights into the phylogeny of flower flies (family Syrphidae). Young, A. D., Skevington, J. H., Mengual, X., Stahls, G., Reemer, M., Jordaens, K., & Lemmon, E. M. (2016). BMC Evolutionary Biology. 16(1), 143.
- Expanding anchored hybrid enrichment to resolve both deep and shallow relationships within the spider tree of life. Hamilton, C.A., Lemmon, A.R., Lemmon, E.M., Bond, J.E. (2016). BMC Evol Biol. 16(1), 212.
- Methodological congruence in phylogenomic analyses with morphological support for teiid lizards (Sauria: Teiidae). Tucker, D.B., Colli, G.R., Giugliano, L.G., Hedges, S.B., Hendry, C.R., Lemmon, E.M., Lemmon, A.R., Sites, J.W. Jr, Pyron, R.A. (2016). Mol Phylogenet Evol. 103, 75-84.
- Nuclear Architecture Organized by Rif1 Underpins the Replication-Timing Program. Foti R, Gnan S, Cornacchia D, Dileep V, Bulut-Karslioglu A, Diehl S, Buness A, Klein FA, Huber W, Johnstone E, Loos R, Bertone P, Gilbert DM, Manke T, Jenuwein T, Buonomo SC. (2016). Mol Cell. 61(2), 260-73.
- Spatio-temporal re-organization of replication foci accompanies replication domain consolidation during human pluripotent stem cell lineage specification. Wilson, K.A., Elefanty, A.G., Stanley, E.G., Gilbert, D.M. (2016). Cell Cycle. 15(18), 2464-75.
- Integrating phylogenomic and morphological data to assess candidate species-delimitation models in Brown and Red-bellied snakes (Storeria). Pyron, R. A., Hendry, C. R., Hsieh, A. R., Lemmon, A. R., & Lemmon, E. M. (2016). Zoological Journal of the Linnean Society. 177(4), 937-949.
- Comprehensive nucleosome mapping of the human genome in cancer progression. Druliner, B. R., Vera, D., Johnson, R., Ruan, X., Apone, L. M., Dimalanta, E. T., & Dennis, J. H. (2016). Oncotarget, 7(12), 13429–45.
- Novel Gefitinib Formulation with Improved Oral Bioavailability in Treatment of A431 Skin Carcinoma. Godugu, C., Doddapaneni, R., Patel, A. R., Singh, R., Mercer, R., & Singh, M. (2016). Pharmaceutical Research, 33(1), 137–54.
- RNA helicase Belle/DDX3 regulates transgene expression in Drosophila. Lo, P.-K., Huang, Y.-C., Poulton, J. S., Leake, N., Palmer, W. H., Vera, D., & Deng, W.-M. (2016). Developmental Biology, 412(1), 57–70.
- Expression Differentiation Is Constrained to Low-Expression Proteins over Ecological Timescales. Margres, M. J., Wray, K. P., Seavy, M., McGivern, J. J., Herrera, N. D., & Rokyta, D. R. (2016). Genetics, 202(1), 273–83.
- The impact of anchored phylogenomics and taxon sampling on phylogenetic inference in narrow-mouthed frogs (Anura, Microhylidae). Peloso, P. L. V., Frost, D. R., Richards, S. J., Rodrigues, M. T., Donnellan, S., Matsui, M., & Wheeler, W. C. (2016). Cladistics, 32(2), 113–140.
- An Examination of Dynamic Gene Expression Changes in the Mouse Brain During Pregnancy and the Postpartum Period. Ray, S., Tzeng, R.-Y., DiCarlo, L. M., Bundy, J. L., Vied, C., Tyson, G., & Arbeitman, M. N. (2016). G3 (Bethesda, Md.), 6(1), 221–33.
- ExtraPEG: A Polyethylene Glycol-Based Method for Enrichment of Extracellular Vesicles. Rider, M. A., Hurwitz, S. N., & Meckes, D. G. (2016). Scientific Reports, 6, 23978.
- Hierarchical regulation of the genome: global changes in nucleosome organization potentiate genome response. Sexton, B. S., Druliner, B. R., Vera, D. L., Avey, D., Zhu, F., & Dennis, J. H. (2016). Oncotarget, 7(6), 6460–75.
- Akt mediated phosphorylation of LARP6; critical step in biosynthesis of type I collagen. Zhang, Y., & Stefanovic, B. (2016). Scientific Reports, 6, 22597.
- Evaluating the performance of anchored hybrid enrichment at the tips of the tree of life: a phylogenetic analysis of Australian Eugongylus group scincid lizards. Brandley, M. C., Bragg, J. G., Singhal, S., Chapple, D. G., Jennings, C. K., Lemmon, A. R., & Moritz, C. (2015). BMC Evolutionary Biology, 15, 62.
- The Anxiolytic and Antidepressant-like Effects of Testosterone and Estrogen in Gonadectomized Male Rats. Carrier, N., Saland, S. K., Duclot, F., He, H., Mercer, R., & Kabbaj, M. (2015). Biological Psychiatry, 78(4), 259–69.
- Topologically associating domains and their long-range contacts are established during early G1 coincident with the establishment of the replication-timing program. Dileep, V., Ay, F., Sima, J., Vera, D. L., Noble, W. S., & Gilbert, D. M. (2015). Genome Research, 25(8), 1104–13.
- The estrous cycle surpasses sex differences in regulating the transcriptome in the rat medial prefrontal cortex and reveals an underlying role of early growth response 1. Duclot, F., & Kabbaj, M. (2015). Genome Biology, 16, 256.
- Are 100 enough? Inferring acanthomorph teleost phylogeny using Anchored Hybrid Enrichment. Eytan, R. I., Evans, B. R., Dornburg, A., Lemmon, A. R., Lemmon, E. M., Wainwright, P. C., & Near, T. J. (2015). BMC Evolutionary Biology, 15, 113.
- Non-adaptive plasticity potentiates rapid adaptive evolution of gene expression in nature. Ghalambor, C. K., Hoke, K. L., Ruell, E. W., Fischer, E. K., Reznick, D. N., & Hughes, K. A. (2015). Nature, 525(7569), 372–5.
- Contrasting modes and tempos of venom expression evolution in two snake species. Margres, M. J., McGivern, J. J., Seavy, M., Wray, K. P., Facente, J., & Rokyta, D. R. (2015). Genetics, 199(1), 165–76.
- Phenotypic integration in the feeding system of the eastern diamondback rattlesnake (Crotalus adamanteus). Margres, M. J., Wray, K. P., Seavy, M., McGivern, J. J., Sanader, D., & Rokyta, D. R. (2015). Molecular Ecology, 24(13), 3405–20.
- In Vivo Analysis of Troponin C Knock-In (A8V) Mice: Evidence that TNNC1 Is a Hypertrophic Cardiomyopathy Susceptibility Gene. Martins, A. S., Parvatiyar, M. S., Feng, H.-Z., Bos, J. M., Gonzalez-Martinez, D., Vukmirovic, M., & Pinto, J. R. (2015). Circulation: Cardiovascular Genetics, 8(5), 653–64.
- A comprehensive phylogeny of birds (Aves) using targeted next-generation DNA sequencing. Prum, R. O., Berv, J. S., Dornburg, A., Field, D. J., Townsend, J. P., Lemmon, E. M., & Lemmon, A. R. (2015). Nature, 526(7574), 569–73.
- Stimulation of the Drosophila immune system alters genome-wide nucleosome occupancy. Ren, Y., Vera, D. L., Hughes, K. A., & Dennis, J. H. (2015). Genomics Data, 3, 146–7.
- Identification of the oncogenic kinase TOPK/PBK as a master mitotic regulator of C2H2 zinc finger proteins. Rizkallah, R., Batsomboon, P., Dudley, G. B., & Hurt, M. M. (2015). Oncotarget, 6(3), 1446–61.
- Post-transcriptional Mechanisms Contribute Little to Phenotypic Variation in Snake Venoms. Rokyta, D. R., Margres, M. J., & Calvin, K. (2015). G3 (Bethesda, Md.), 5(11), 2375–82.
- The transcriptomic and proteomic basis for the evolution of a novel venom phenotype within the Timber Rattlesnake (Crotalus horridus). Rokyta, D. R., Wray, K. P., McGivern, J. J., & Margres, M. J. (2015). Toxicon, 98, 34–48.
- Comparing species tree estimation with large anchored phylogenomic and small Sanger-sequenced molecular datasets: an empirical study on Malagasy pseudoxyrhophiine snakes. Ruane, S., Raxworthy, C. J., Lemmon, A. R., Lemmon, E. M., & Burbrink, F. T. (2015). BMC Evolutionary Biology, 15, 221.
- Methyl supplementation attenuates cocaine-seeking behaviors and cocaine-induced c-Fos activation in a DNA methylation-dependent manner. Wright, K. N., Hollis, F., Duclot, F., Dossat, A. M., Strong, C. E., Francis, T. C., & Kabbaj, M. (2015). The Journal of Neuroscience, 35(23), 8948–58.
- Differential effects of acellular embryonic matrices on pluripotent stem cell expansion and neural differentiation. Yan, Y., Martin, L. M., Bosco, D. B., Bundy, J. L., Nowakowski, R. S., Sang, Q.-X. A., & Li, Y. (2015). Biomaterials, 73, 231–42.
- The genome, proteome and phylogenetic analysis of Sinorhizobium meliloti phage ΦM12, the founder of a new group of T4-superfamily phages. Brewer, T. E., Stroupe, M. E., & Jones, K. M. (2014). Virology, 450-451, 84–97.
- Phenotypic and genomic plasticity of alternative male reproductive tactics in sailfin mollies. Fraser, B. A., Janowitz, I., Thairu, M., Travis, J., & Hughes, K. A. (2014). Proceedings. Biological Sciences / The Royal Society, 281(1781), 20132310.
- A hybrid phylogenetic-phylogenomic approach for species tree estimation in African Agama lizards with applications to biogeography, character evolution, and diversification. Leaché, A. D., Wagner, P., Linkem, C. W., Böhme, W., Papenfuss, T. J., Chong, R. A., & Burbrink, F. T. (2014). Molecular Phylogenetics and Evolution, 79, 215–30.
- Linking the transcriptome and proteome to characterize the venom of the eastern diamondback rattlesnake (Crotalus adamanteus). Margres, M. J., McGivern, J. J., Wray, K. P., Seavy, M., Calvin, K., & Rokyta, D. R. (2014). Journal of Proteomics, 96, 145–58.
- RNA-seq and high-definition mass spectrometry reveal the complex and divergent venoms of two rear-fanged colubrid snakes. McGivern, J. J., Wray, K. P., Margres, M. J., Couch, M. E., Mackessy, S. P., & Rokyta, D. R. (2014). BMC Genomics, 15, 1061.
- Effectiveness of phylogenomic data and coalescent species-tree methods for resolving difficult nodes in the phylogeny of advanced snakes (Serpentes: Caenophidia). Pyron, R. A., Hendry, C. R., Chou, V. M., Lemmon, E. M., Lemmon, A. R., & Burbrink, F. T. (2014). Molecular Phylogenetics and Evolution, 81, 221–31.
- Hydrogen-Deuterium Exchange Mass Spectrometry of Phosphorylation Induced Changes in Smooth Muscle Myosin. Sen, K. I., Haldeman, B. D., Brizendine, R., Mercer, R., Cremo, C. R., & Fajer, P. G. (2013). Biophysical Journal, 104(2), 452a.
- Anchored hybrid enrichment for massively high-throughput phylogenomics. Lemmon, A. R., Emme, S. A., & Lemmon, E. M. (2012). Systematic Biology, 61(5), 727–44.
- High-throughput identification of informative nuclear loci for shallow-scale phylogenetics and phylogeography. Lemmon, A. R., & Lemmon, E. M. (2012). Systematic Biology, 61(5), 745–61.
- Repli-seq: genome-wide analysis of replication timing by next-generation sequencing. Marchal, C., Sasaki, T., Vera, D., Wilson, K., Sima, J., Rivera-Mulia, J-C., Trevilla Garcia, C., Nogues, C., Nafie, E., Gilbert, D.M. (2017). In Review.


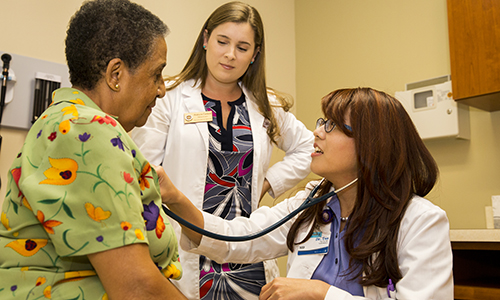 Every year, thousands of new students arrive at Florida State hoping to enter a career in health care. For a variety of reasons, most of these would-be physicians, nurses, pharmacists and others wind up in different careers — and communities lose out on the health care they could have provided.
Every year, thousands of new students arrive at Florida State hoping to enter a career in health care. For a variety of reasons, most of these would-be physicians, nurses, pharmacists and others wind up in different careers — and communities lose out on the health care they could have provided. TALLAHASSEE, Fla. — Three years after discovering that a single, unidentified mechanism was modifying about 800 proteins simultaneously during cell division, Florida State University researchers have identified that mystery enzyme.
TALLAHASSEE, Fla. — Three years after discovering that a single, unidentified mechanism was modifying about 800 proteins simultaneously during cell division, Florida State University researchers have identified that mystery enzyme. Scientists have come a long way since creating the first map of the human genome in 2003. Technological advances have accelerated DNA sequencing of the human genome – a process that once took a decade or more to complete – to the point that it can be done in a matter of days, even while yielding more information.
Scientists have come a long way since creating the first map of the human genome in 2003. Technological advances have accelerated DNA sequencing of the human genome – a process that once took a decade or more to complete – to the point that it can be done in a matter of days, even while yielding more information. For most people, the word “research” conjures up images of serious-looking scientists in white coats, beakers filled with colorful liquids, microscopes and cluttered laboratories.
For most people, the word “research” conjures up images of serious-looking scientists in white coats, beakers filled with colorful liquids, microscopes and cluttered laboratories. Dr. Roger Mercer has been named director of the College of Medicine Translational Science Laboratory. Dr. Mercer will be responsible for the ongoing development, overall operation and coordination of daily technical activities of a new state-of-the-art facility for the identification of biomarkers taken from biological samples and for high end proteomic and metabolomic analyses. With more than 25 years experience in academia and in the biotech industry, Dr Mercer will play a leadership role in the College of Medicine’s Translational Science Initiative within its multi-site clinical research network.
Dr. Roger Mercer has been named director of the College of Medicine Translational Science Laboratory. Dr. Mercer will be responsible for the ongoing development, overall operation and coordination of daily technical activities of a new state-of-the-art facility for the identification of biomarkers taken from biological samples and for high end proteomic and metabolomic analyses. With more than 25 years experience in academia and in the biotech industry, Dr Mercer will play a leadership role in the College of Medicine’s Translational Science Initiative within its multi-site clinical research network. An ordinarily quiet corner of the College of Medicine’s Research Building was teeming with science-minded sightseers Aug. 25 for the open house of the Translational Science Laboratory.
An ordinarily quiet corner of the College of Medicine’s Research Building was teeming with science-minded sightseers Aug. 25 for the open house of the Translational Science Laboratory.




